Research
Post-transcriptional regulation is a key stage of gene regulation and comprises the homeostasis of mRNA levels for a functional cell. Misbalance of these levels is a major cause for diseases like autoimmunity and cancer.
We are studying regulatory RNA-protein complexes that decide about the fate of mRNAs and thereby form a major regulatory parameter in cell development, during immune responses and in cancer progression. mRNAs contain untranslated regions (UTRs) in their 5’ and 3’ termini that are full of cis-regulatory elements which can either be structured moieties like stem-loops, extended double-stranded stretches of RNA or short linear sequences. It appears that mRNAs are regulated through the interplay of multiple of these cis elements, that are eventually organized in hubs. trans-acting factors are mainly microRNAs and RNA-binding proteins (RBPs) with their RNA-binding domains (RBDs) that recognize the cis elements and provoke downstream effects decisive for the fate of the mRNA. These can result in mRNA decay, protection, splicing, transport or its translational inhibition. Apparently the engagement of mRNA and proteins is a key checkpoint for the regulation of mRNA turnover and is not only affected by the expression levels of RBPs, but also by the spatiotemporal availability and organization of relevant RNA cis elements. Those might be masked in RNA secondary structure, by modifications or other RBPs.
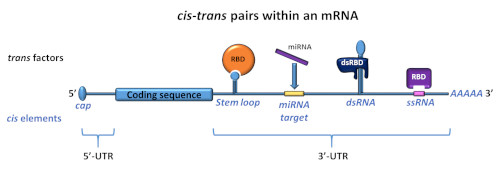
Scheme of an mRNA with various cis-regulatory elements in its 3’-untranslated region (UTR). These are recognized by trans factors like RNA-binding proteins through cognate RNA-binding domains (RBDs).
Nature has evolved various concepts of RNA-protein interactions that we are only starting to understand in detail. RBPs normally find their targets either through a highly-specific and affine recognition of structured RNA cis elements (e.g. the by the Roquin ROQ domain) or the multivalent interaction of multiple RNA-binding domains (RBDs) with sequentially arranged short linear single-stranded cis elements (e.g. as known for IMP proteins). In either case, the prediction and identification of the exact target mRNAs is hampered by a likely higher-order organization of extended RNA regions involved in the interaction with the protein. The more, a certain degree of plasticity is achieved by complex-intrinsic dynamics and molecular flexibility.
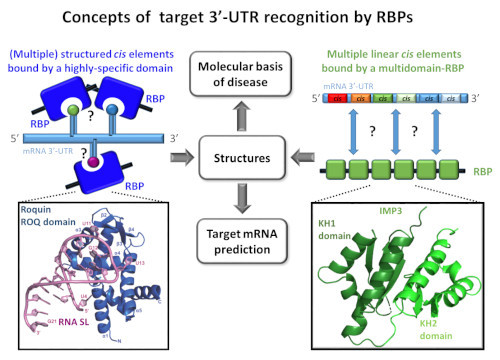
Two concepts of mRNA 3’-UTR recognition by either a highly specific domain binding to structured RNA (left, exemplified by the Roquin ROQ domain bound to a stem-loop RNA element) or a multidomain RBP that encounters sequential short linear RNA elements (right, a tandem KH domain of IMP3 represents two of six potential RBDs), both to be found in disease-related mRNA regulation.
In order to dissect the networks of regulatory cis-trans interactions we determine atom-resolved pictures of mRNA 3’-UTR cis-regulatory elements in complex with RBPs using a broad set of methods from structural biology complemented by RNA- and protein-biochemical/biophysical methods. We are working in close collaboration with scientists from the fields of cell biology, immunology and big data for the mutual exchange of new finding and the transfer of identified structural determinants into living systems. We thereby hope to contribute to a broader understanding of mRNA-encoded gene regulation at the onset of diseases.
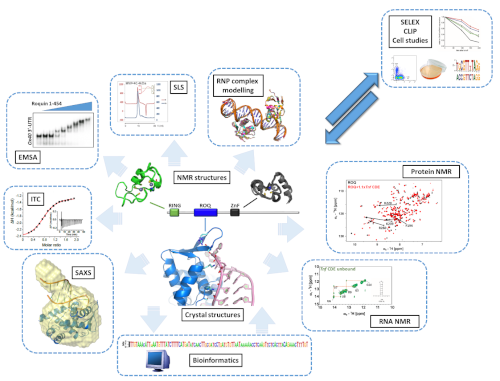
Our Integrated Structural Biology approach implements all kinds of methods to help obtaining molecular pictures and derive mechanistic concepts that are later on validated in functional assays with collaborating groups. In turn, we also exploit data that suggest novel target mRNAs and their cis elements to be regulated by our RBPs of interest.

Dr. Andreas Schlundt
Biozentrum, Campus Riedberg
Building N250, Room 101
Max-von-Laue-Str. 9
60438 Frankfurt am Main
T +49 69 798-29699
F +49 69 798-29527
E schlundt@bio.uni-frankfurt.de
- Studying at Goethe University
- International applicants
- Faculties
- Overview of study programmes
- Programme for refugees
- GRADE
- Goethe Business School (continuing education)
- Research at Goethe University
- Scientific news
- Goethe Welcome Center (for international researchers)
- Collaborative research projects
- Individual research
- Visiting fellowships
- Endowed chairs
- About the University
- News-in-brief
- University administration
- Campus locations
- Campus life
- University archives (German)
- Rhine-Main-Universities





