Projects

Framework and questions:
- Project designed for 3 years
- How can students get involved in contributing to sustainability?
- How can we learn from each other sustainably?
- What contribution can we make to sustainability at Goethe University?
- Where are our own processes in need of improvement in terms of sustainability?
- What can we do more efficiently?
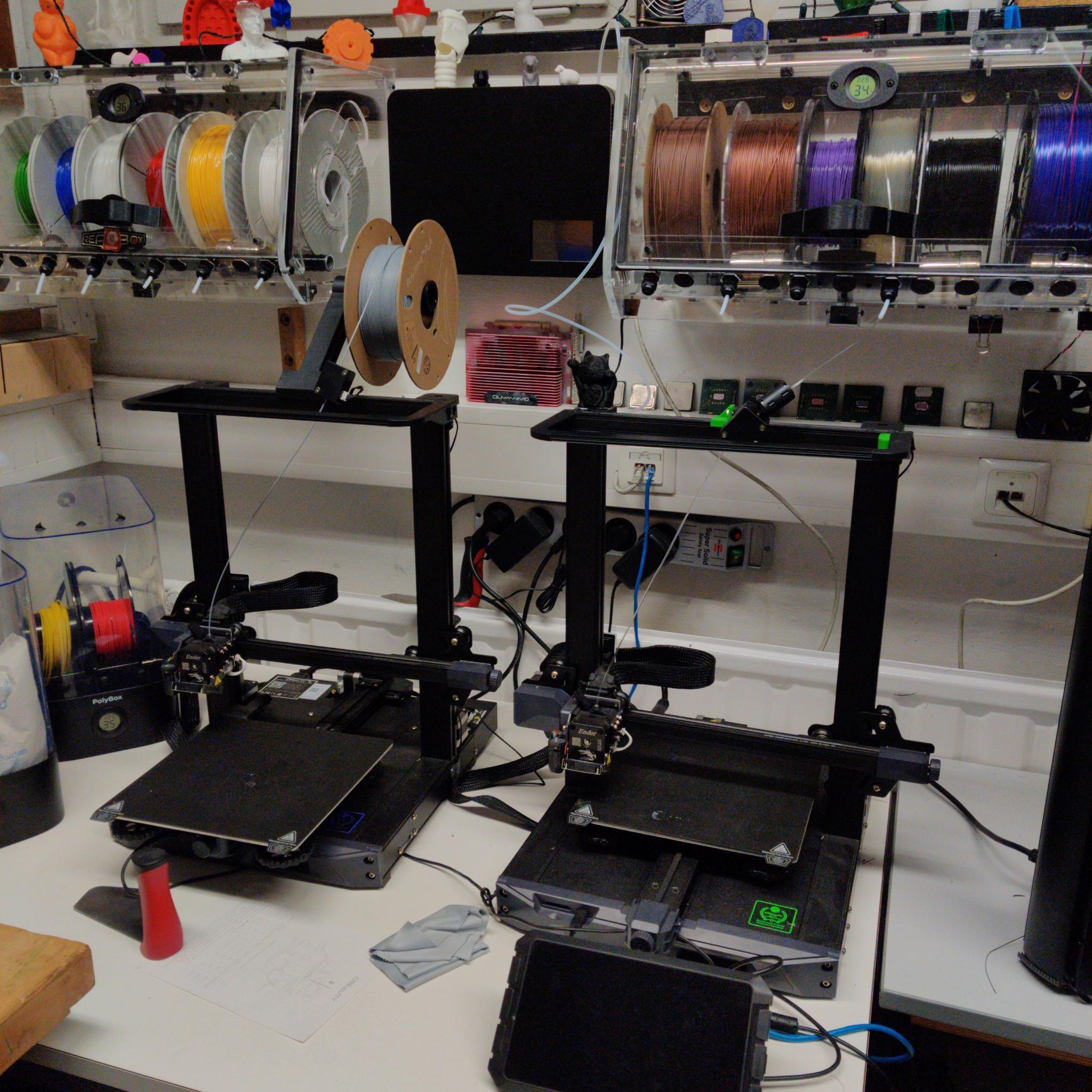
Measures to date include:
- Material-friendly storage and quality-preserving measures in the FDM printing process
- Material preparation & recovery in the SLS printing process
- Sustainability in the project and university
- Example project SimonsVoss Transponder Sleeve
- Recycling of printing waste
- Energy-saving measures, e.g. through
- Remote access 3D printer & power supply
- Optimization of printer profiles


At our Maker Cafés, which take place several times a year, students can come together in a cozy atmosphere and get to know the possibilities of our Makerspace. There are usually one ore more workshops in various areas.


Partnership Accelerate Rapid Prototyping And Repair Culture (ARPARC) is taking place as part of the „Digital Teaching and Learning Lab“.





Framework and questions:
- Project designed for 3 years
- How can students get involved in contributing to sustainability?
- How can we learn from each other sustainably?
- What contribution can we make to sustainability at Goethe University?
- Where are our own processes in need of improvement in terms of sustainability?
- What can we do more efficiently?
Measures to date include:
- Material-friendly storage and quality-preserving measures in the FDM printing process
- Material preparation & recovery in the SLS printing process
- Sustainability in the project and university
- Example project SimonsVoss Transponder Sleeve
- Recycling of printing waste
- Energy-saving measures, e.g. through
- Remote access 3D printer & power supply
- Optimization of printer profiles


At our Maker Cafés, which take place several times a year, students can come together in a cozy atmosphere and get to know the possibilities of our Makerspace. There are usually one ore more workshops in various areas.


Partnership Accelerate Rapid Prototyping And Repair Culture (ARPARC) is taking place as part of the „Digital Teaching and Learning Lab“.




This 3D DNA model was printed and constructed at MakeLab as part of Critical Gemonics. Critical Genomics is a student initiative at Goethe University that organizes an interdisciplinary lecture series and a summer school on genomics. It will take place from April to August 2024 in Frankurt am Main. Its aim is to create a time and place where students from the humanities and life sciences can educate each other through critical discussion about the field of genomics.
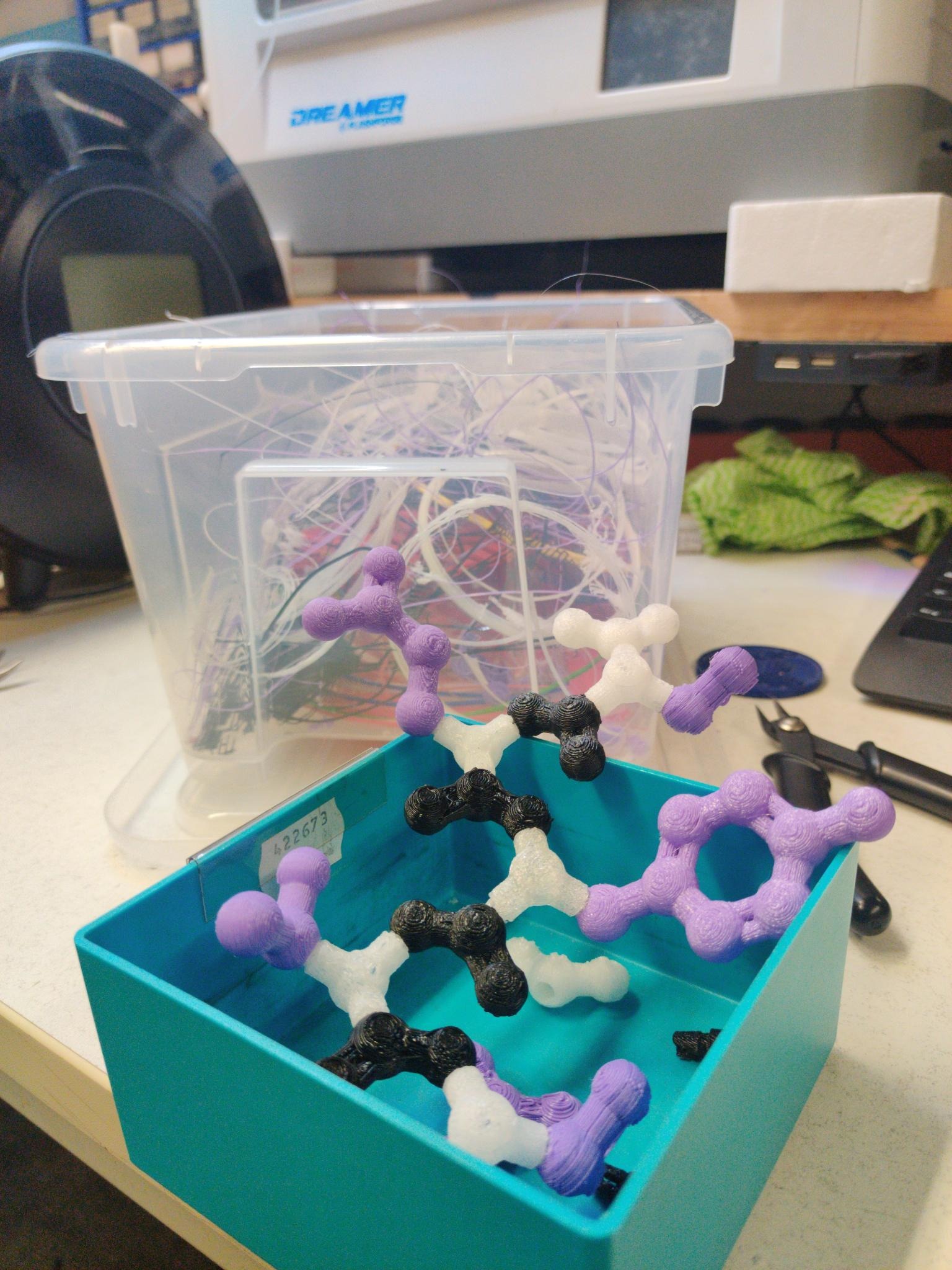


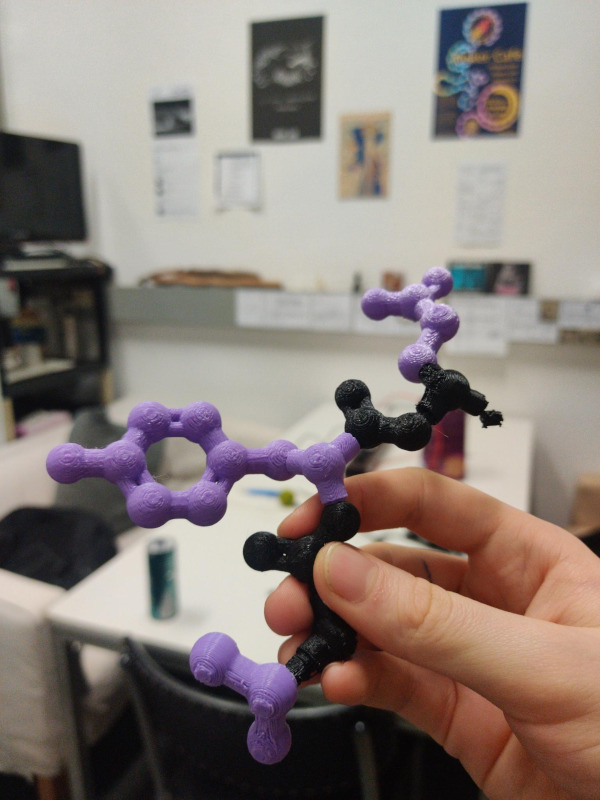
For a project by bioinformatics students, 3D-printed amino acid models were created to visualize the composition of protein chains and to practice these with the modular models.

This 3D DNA model was printed and constructed at MakeLab as part of Critical Gemonics. Critical Genomics is a student initiative at Goethe University that organizes an interdisciplinary lecture series and a summer school on genomics. It will take place from April to August 2024 in Frankurt am Main. Its aim is to create a time and place where students from the humanities and life sciences can educate each other through critical discussion about the field of genomics.


For a project by bioinformatics students, 3D-printed amino acid models were created to visualize the composition of protein chains and to practice these with the modular models.

- Studying at Goethe University
- International applicants
- Faculties
- Overview of study programmes
- Programme for refugees
- GRADE
- Goethe Business School (continuing education)
- Research at Goethe University
- Scientific news
- Goethe Welcome Center (for international researchers)
- Collaborative research projects
- Individual research
- Visiting fellowships
- Endowed chairs
- About the University
- News-in-brief
- University administration
- Campus locations
- Campus life
- University archives (German)
- Rhine-Main-Universities